This is a major release that fixes several bugs and incorporates many new features on NMR, MS, the Molecule Editor and the GUI and this new version also includes the release of the Mnova Screen plugin, an automatic tool for the analysis of ligand screening NMR data.
-
New features
Capability to automatically setup 1D spectra on the projections of a 2D
There is a new preference (Auto Attach Traces) under the menu ‘Edit/Preferences/NMR/Import’ to automatically use the 1D spectra as external traces of a 2D.

Assignments made on a 2D spectrum are transferred to the multiplets in 1H spectrum
Once an assignment is made in a 2D spectrum, this assignment will be transferred to the applicable multiplet in the 1H spectrum.

New script to report the assignments into a table
Assignments reports can be generated via Scripts/Report/Assignments. It is required at least a 1H spectrum. The table will only include columns for the spectra present in the document containing assignments. It generates a table in which each row corresponds to each non-equivalent proton. Including 2D correlations is optional from Setup Assignment Report dialog, and it may also show the chemical shift. When no chemical shift is included, the correlations get ordered according to the atom number. When chemical shifts are included, the correlations get ordered according to the values of chemical shifts.
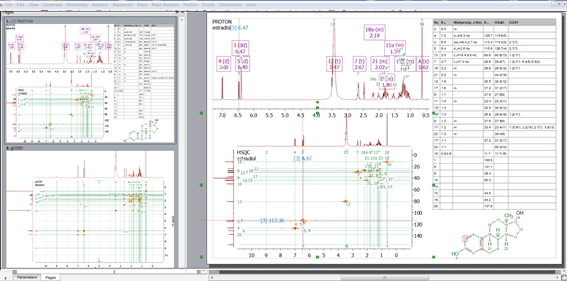
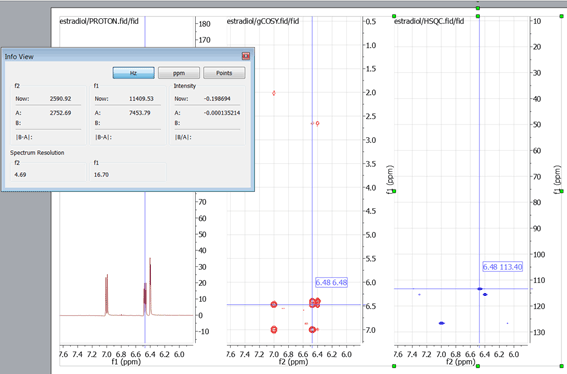
Synchronized crosshair (vertical and horizontal) in COSY spectra when used on top of HSQC spectra
When hovering the crosshair over the HSQC spectrum, vertical and horizontal lines are displayed in the COSY spectrum.
Layout template remembers the zoom ranges
It will be possible to create layout templates keeping the spectral widths.
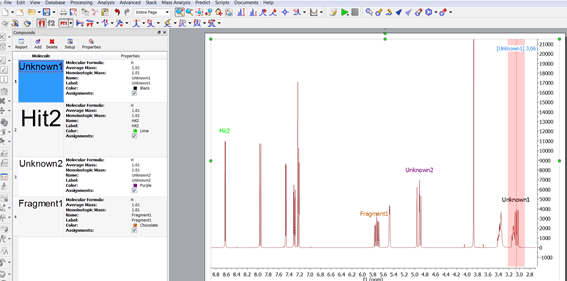
Assignment labels of Unknown compounds are showed with the Compound color
It is possible to select different colors for the assignments of unknown compounds
Atom renumbering changes assignment label automatically on peaks and multiplets
When changing the atomNumber, the assignment label is automatically updated in the peaks and multiplets labels shown in the spectrum.
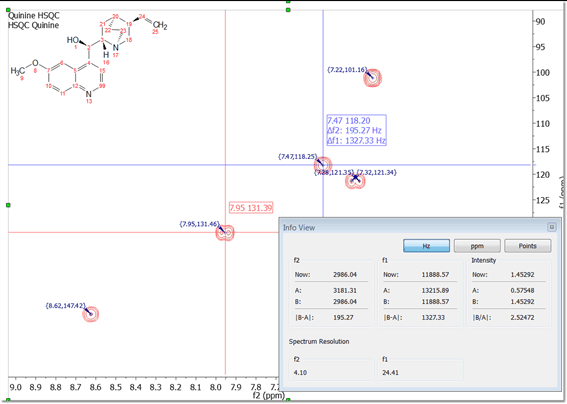
Differences and shifts of lines (crosshair) flying with the crosshairs
When using the crosshair, information about position (in the units selected for the spectrum scales) and differences (in Hz) are flying close to the cursor.
Bugs fixed.
- Spectra from 1r with ZF get distorted when superimposed and auto scaled
- Problems when setting vertical scale as logarithmic in Data Analysis graph
- Peaks and Multiplets misaligned after opening and canceling the processing template
- Residual curves do not show for some peaks
- Problems when loading directories containing mrc files
- Duplicated assignment labels after changing the atom label
- Redo after editing a multiplet or integral did not work
- Assignment labels are not updated when the user writes the new atom Number directly over an atom.
- Wrong behavior when zooming on graph items
- Difference spectrum resulting from Arithmetic tool showed out of phase
- Align Spectra: preview did not work
- Problems selecting 30 spectra and opening Arithmetic tool
- Problems hovering the mouse over a molecule which has been modified in another document
- Problems disabling points in Data Analysis using the scissors tool
- Wrong options in Normalize dialogs when normalizing by Total Area or Manual
- Wrong binning result after rescaling in binned stacked spectra
- Wrong automatic phase correction in F2 of a HSQC from Varian
- Carbon dimension out of phase in gradient HMBC spectra from Varian
- HSQC from Varian showed out of phase when loaded in Mnova
- Export Fit Regions script did not export all the regions of all spectra in a stack
- Error from Multiplet Report to Spectrum script when parsing a - sign
- Wrong Integral list when saving as 1D Integral Series for stacked spectra with visual decimation
- Gamma value not read in a 7Li DOSY spectrum
- Transitions were shifted a little bit with respect to the spectrum
- When loading 1r files the title did not show the folder in which the spectrum is located
- Temperature was not correctly read from JCAMP-DX
- Problems when adding a peak of a Line Fitting region in a different page
- Duplicated assignment labels after changing the atom label
- Problems when applying binning to a range
- Problems opening PCA panel having 36 spectra overlaid
- Average Window parameter shows now as "RMS calculation span (points)"
- Multiple chemical shift labels (from assignments) on a molecule
- Multiplet spectrum parser failed with special characters
-
New features
Ability to show molecule annotation info below molecule. The molecule contains a label (available in Molecule Properties/General) which can be customized to include free text and also several parameters (as shown in the figure below).
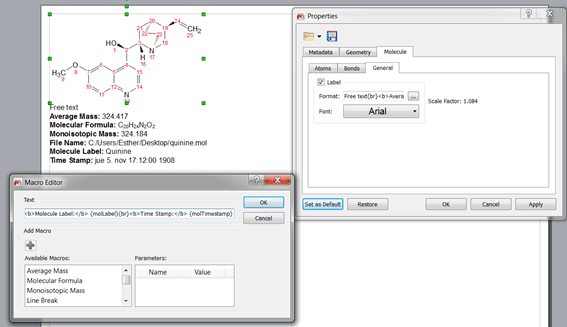
A quick way of renumbering molecules in a mixture to avoid repeated numbers
Clicking the 'Concatenate Numbering' button, will de-duplicate the atom numbers of all the molecules in the document. The molecules will be renumbered so that the atom numbers are not repeated (leaving a gap from one molecule to another). The order in which molecules get renumbered will depend on the size of the molecule (the biggest molecule will keep its numbering).

-
New features
Capability to add a parameter in the title as a macro via Mass Spectrum Properties
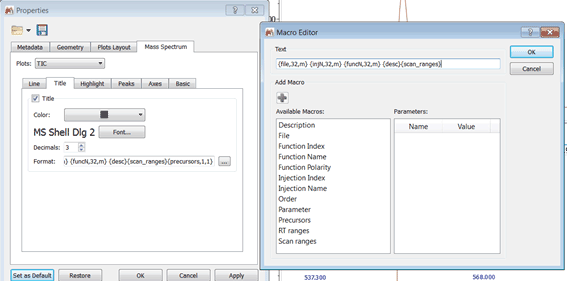
MS Browser: double click to open a trace
Double clicking on the applicable trace from the MS Browser will automatically display the trace in the spectral window
Bugs Fixed
- Deleted TIC from top gets restored at the bottom when making Undo
- Not possible to select two spectra and co-add them
-
New Data Browser
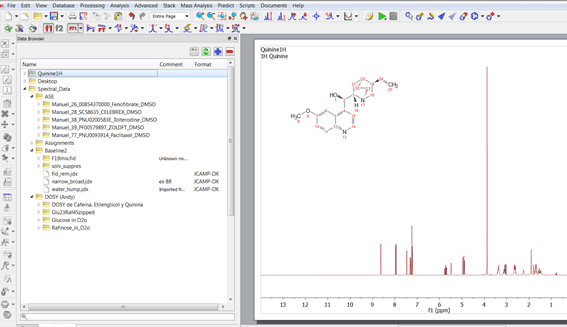 A new tree view data browser has been implemented with the following features: 1.Capability to register several data locations just by clicking on the applicable button or by dragging and dropping the folder containing the datasets.
A new tree view data browser has been implemented with the following features: 1.Capability to register several data locations just by clicking on the applicable button or by dragging and dropping the folder containing the datasets.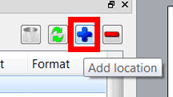 These data locations are the same as in "Preferences>General>Open&Save>Open Locations". 2.The data browser will handle all spectral and molecule formats from all vendors, but the user will be entitled to filter some of the formats:
These data locations are the same as in "Preferences>General>Open&Save>Open Locations". 2.The data browser will handle all spectral and molecule formats from all vendors, but the user will be entitled to filter some of the formats: 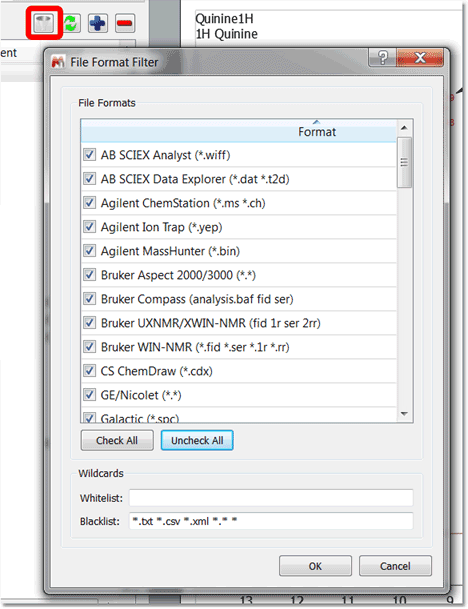 3.In case of Bruker datasets all openable data will be listed in the tree view: fid and all 1r's or ser and all 2rr's. 4.The data folders are sorted numerically. 5.Spectral titles will be shown directly in the panel and also be available as tool-tips. 6.Double click to open datasets. It will be possible to tick several datasets to open. Hovering the cursor over the spectra folder or fid/ser file related will display some information. In case of Bruker files, the parm.txt file is shown as tooltip. In case of Varian files, the content of Comment field will be displayed.
3.In case of Bruker datasets all openable data will be listed in the tree view: fid and all 1r's or ser and all 2rr's. 4.The data folders are sorted numerically. 5.Spectral titles will be shown directly in the panel and also be available as tool-tips. 6.Double click to open datasets. It will be possible to tick several datasets to open. Hovering the cursor over the spectra folder or fid/ser file related will display some information. In case of Bruker files, the parm.txt file is shown as tooltip. In case of Varian files, the content of Comment field will be displayed.- If the panel is empty, it will show a message inside the panel: "Add your dataset locations here"
- The search feature works with wildcards and strings in the same way. For example, you will get the same result by typing ".mol" or "*.mol" in the search cell, and both searches take equal time:
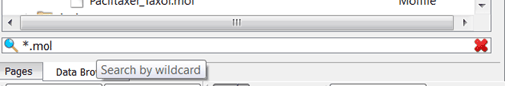
Drag and drop spectrum name opens all spectra below spectrum name in the order of the numbers
If you have a main folder, containing several sub-folders with different experiments, it is now enough to drag&drop the main folder into Mnova in order to load all the experiments below (sorting them by numbering).
Vertical Tabs in Dock Areas
It will display the tabs vertically in the dock widgets (useful when having a lot of dock widgets opened in the lateral dock and most of the tabs displayed horizontally are not visible):
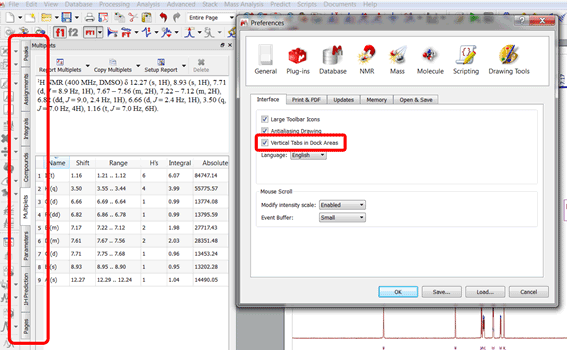
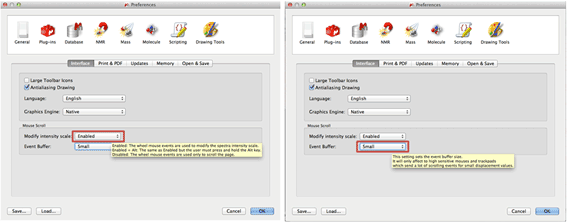
New settings for magic mouses and trackpads
The 'Modify Intensity Scale' option enables/disables changing the spectra intensity/threshold via the mouse wheel. It also allows to enable it but requiring that the Alt key is pressed. The 'Event Buffer' will only affect to high sensitive mouses/trackpads (Like the magic mouse and magic trackpad from Mac) in any other device the change of this setting won't affect. It sets the event buffer size (or even disable it).
New features in the scripting engine
Function to align non stacked spectra: AlignFunction() Capability to Print from a script: mainWindow.activeDocument.print();
Bugs Fixed
- Problems when deleting a compound from Compounds table with Enable Undo option off
- Problems trying to show a molecule with Ir atom in the 3D Molecule panel
- Problems under Linux when painting the Molecule after removing Explicit Hydrogens
- Molecule did not get restored correctly after resizing and clicking Undo
- Atom numbers could not be blank
- Problems report all the compound properties
- Problems using isotope information to calculate mass information
-
Bugs Fixed
- Avoid duplicated labels in a Simulation when editing the group labels
- Problems when trying to getting a spin system from multiplets
- Problems loading a spin simulation xml file containing different groups with the same name
- HSQC predicted showed wrong
- Batch prediction did not predict more than 32K mols
- Editing the predicted spectrum from the Spin Simulation table was not working
- Problems with the line width in predicted and simulated spectra
- Spin Simulation: The changes in the line width value were not applied to the predicted spectrum
- Highlighting was not working fine in the Spin Simulation Panel
- Horizontal and vertical panning did not work when only NMR Predict Desktop was licensed
- Numbering gets changed when updating a Prediction DB from a sdf file
-
Bugs Fixed
- Assignments were not shown after verification
- Verification Results showed in gray after running Verification
- Verification failed with 'perch' molecules
-
Bugs fixed:
- Error when trying to save a record and NMR plugin was not installed


