A versatile, functional program for the quantitation of mixture components by NMR based on chemical shift range integrals.
SMA (Simple Mixtures Analysis by NMR) is only “simple” by name!
This advanced solution from Mnova provides a comprehensive workflow to perform targeted analyses of mixtures with the aim of determining the presence and quantity of a given component. It allows screening for impurities, degradants, adulterants, etc., in consumer goods and in raw material, and reports on their concentrations. Mnova SMA is hugely flexible and customizable, and can be easily implemented as a semi- or fully automated solution depending on your needs. Check out our fully automated solution at Mnova Gears SMA.
A valid license of Mnova NMR is required to use Mnova SMA.


“Roll your own” targeted mixtures analysis!
Mnova SMA: 45-day FREE trial
 1. Download
1. Download
SMA is a plugin integrated in Mnova. No extra installer is required.
 2. Installation
2. Installation
Unzip the file and run it. You’ll be redirected to our website to request a license number.
 3. License
3. License
Fill in the license request form and you will receive your trial license via e-mail.
SMA
Features
There is an almost endless list of applications where SMA will automatically carry out your mixtures analysis for you.
Define the targeted analysis application using the formula editor to be in full control of the quantification.
Recent high-profile cases have brought attention to the need for goodanalytical procedures.
QC and production environments can benefit from automating workflows using SMA.
Design your own experiments, select your mixture components, include/exclude multiplet ranges, set up integration options and formulae, choose colors for display, personalize the alerts, etc and save your experiment in a library for further use and re-use in your organization.
Customize your reports to include the information you need and choose the preferred formats (PDF, MNOVA, HTML, XML, CSV) and layouts.
Import chemical shift ranges, number of nuclides and molecular weight from your Mnova DB and build your SMA experiment with a few buttons clicks!
In addition, when a standard analysis such as concentration or purity determination is required for quantifying several mixtures, SMA can be quickly put to work: a mixture spectrum can be searched against a DB with spectra of potential components to help identify potential components. Potential hits are reported, and the list can be easily refined by a quick visual inspection.
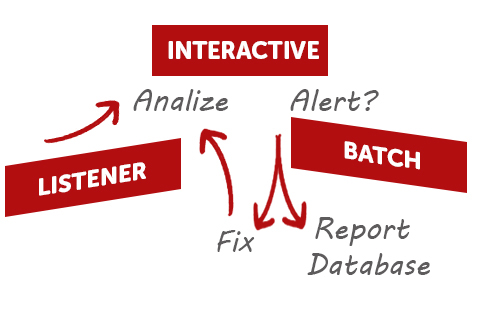
Optimize your workflow and run SMA analyses in automation. Gears SMA will process your raw data in Batches from a specified directory or in Real Time as they are produced by the spectrometer.
Your business will certainly appreciate it!
Related applications
Mixture analyses with Mnova
Mnova offers multiple ways to address the analyses of mixtures when presented in NMR spectra, whether they are targeted or untargeted, and whether they require component identification and/or quantification. The most suitable technique depends upon the application.
Discover which solution is best for your needs! And Contact us for more information.
SMA Users
Markets
- Foods, forensics, nutraceuticals, cosmetics, body fluids, impurity levels in fine, chemicals (including solvents), edible oils, wines and fortified alcoholic drinks, fruit juices.
- QC & production environments (batch and real time analysis)




Frequently Asked Questions
FAQS
The only basic requirement is to have a component with at least one peak area in a defined region of a spectrum and to know the number of nuclei which generate the peak. The SMA plugin will use the integral value of this peak to quantify the species in mixtures with a discrete number of components.
SMA is a workflow designed for targeted screening and quantification of components in mixtures. It allows you to determine the concentration of various components simultaneously and is therefore useful when analyzing mixtures of medium complexity, or when determining the purity of samples which are known to regularly contain high levels of impurity that would never be considered as ‘pure’ samples.
SMA’s workflow suits regulated environments where the same compounds and analytical conditions are repeated, and fine control over the analysis is needed.
An automated batch or Real time version of SMA is also useful when medium to high throughput and minimal operator intervention is required (Gears SMA).
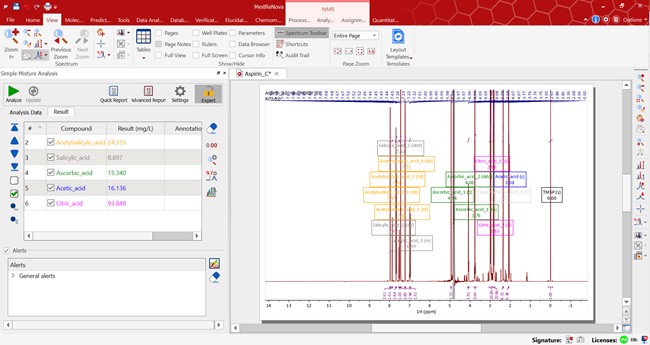
When you hit the ‘Analyze button’, peak picking is applied and multiplets are determined for each region. Then components are quantified using a predefined mixture analysis which converts the integrals to concentrations according to the specified equation. Each component is evaluated in turn and any errors are hidden. Finally, customized results are displayed in the results table and mixture components are highlighted on the spectrum.
SMA is a versatile tool which allows you to specify the formula for your calculations. Integral values are converted into relevant concentrations using mathematical expressions. SMA has all the standard equations for determining Purity, Mass% and Concentration. It’s also easy to define any other equation for concentration by creating a User Defined Equation. For more information about that see our blog post.
- For simple molar ratios between two components you can use the following the general expression:

- If for instance you want to estimate the concentration of your mixtures components, then the concentration is determined using the simple equation:

The Concentration Conversion Factor (CCF) is like a “response factor” for NMR that converts the absolute integral/nuclide to concentration. For more information about the CCF factor please see Q: “What exactly is the CCF factor?”
- Alternatively if you are aiming to calculate the purity of your mixture components, once you create at least one reference material, the fundamental equation for purity (see below) can be applied.

Yes, you have full control of your quantification method specifying the integral regions to be used, and using the equation editor.SMA offers you the flexibility to define the way you quantify your mixtures.These integrals are converted into relevant concentrations using user-defined mathematical expressions. Sample-specific data (weights, etc.) may be required depending on the quantification method used: , Purity%, Mole%, Concentration, etc. See Q: What sort of calculation does SMA use for quantification?
The Peak Pattern Recognition (PPR) is one of Mestrelab’s tools for multiplet extraction from regions where peaks overlap. It uses a spectrum of the pure compound as a reference to find multiplet patterns in a user-defined search window. The tool searches for all peaks within this window in the mixture spectrum and discards those whose integral is greater than a pre-set threshold. Next, the Algorithm tool is called to find the reference spectrum peaks in the set of peaks in the mixture. If found, SMA creates a multiplet and prints (alert window) the probability.
SMA is a versatile tool which can optimize your production process. It can be easily implemented in your workflow for single, batch, or Real time mixture analyses. Methods/SOPs can be saved and used repetitively for consistent results and can be reviewed and updated by experts when required. Methods for as many analyses as needed can be created. Results and reports can be customized. An internal database can be created. Periodic checks of procedures and results will be required for compliance with standards (GxP).
SMA enables you to create a fully automated analytical workflow. That means that spectral processing, peak picking, and integration can all be automated. SMA makes use of all the amazing Mnova features, such as processing templates, and accesses all the peak picking and integration options available in the Mnova NMR plugin, including Global Spectral Deconvolution (GSD) which automatically detects and identifies solvents and low-level impurities.

CCF stands for Concentration Conversion Factor or the Spectrometer Factor. Basically, when the area of a multiplet is determined, this is related to its concentration
C ∝ AI/NN (1)
Where C is the concentration of a chemical species, AI is the absolute integral for a multiplet attributable to that species, and NN is the number of nuclides (Hs, usually) for that multiplet. From this we derive:
C= CCF * AI/NN (2)
A proportionality constant which we call the CCF, allows us to convert a measured AI/NN into a concentration. Fundamentally, the CCF only needs to be determined once for an experiment, and then it is applicable to all chemical species in the reaction! Once that has been performed, all other species concentrations can be determined by applying equation (2), where the AI is determined by the software, and NN is provided by the user. For further information about how to calculate the CCF factor you can read the following qNMR_CCF document.
Type your formula in the Formula editor then use the automatic Check to see if your formula is correct. In the Formula editor you have legends and some additional options to facilitate your task. You can read more about this in our blog post.
Yes, you can run SMA in automation with Mnova Gears (check Gears SMA). This plugin will allow you to process and analyze data files in batches or in real time as the data comes off the spectrometer, and generate reports that are automatically saved to the directory of your choice on your disks or even in the Mnova DB.